Authors:
(1) Shadab Ahamed, University of British Columbia, Vancouver, BC, Canada, BC Cancer Research Institute, Vancouver, BC, Canada. He was also a Mitacs Accelerate Fellow (May 2022 - April 2023) with Microsoft AI for Good Lab, Redmond, WA, USA (e-mail: shadabahamed1996@gmail.com);
(2) Yixi Xu, Microsoft AI for Good Lab, Redmond, WA, USA;
(3) Claire Gowdy, BC Children’s Hospital, Vancouver, BC, Canada;
(4) Joo H. O, St. Mary’s Hospital, Seoul, Republic of Korea;
(5) Ingrid Bloise, BC Cancer, Vancouver, BC, Canada;
(6) Don Wilson, BC Cancer, Vancouver, BC, Canada;
(7) Patrick Martineau, BC Cancer, Vancouver, BC, Canada;
(8) Franc¸ois Benard, BC Cancer, Vancouver, BC, Canada;
(9) Fereshteh Yousefirizi, BC Cancer Research Institute, Vancouver, BC, Canada;
(10) Rahul Dodhia, Microsoft AI for Good Lab, Redmond, WA, USA;
(11) Juan M. Lavista, Microsoft AI for Good Lab, Redmond, WA, USA;
(12) William B. Weeks, Microsoft AI for Good Lab, Redmond, WA, USA;
(13) Carlos F. Uribe, BC Cancer Research Institute, Vancouver, BC, Canada, and University of British Columbia, Vancouver, BC, Canada;
(14) Arman Rahmim, BC Cancer Research Institute, Vancouver, BC, Canada, and University of British Columbia, Vancouver, BC, Canada.
Table of Links
Conclusion and References
III. MATERIALS AND METHODS
A. Dataset
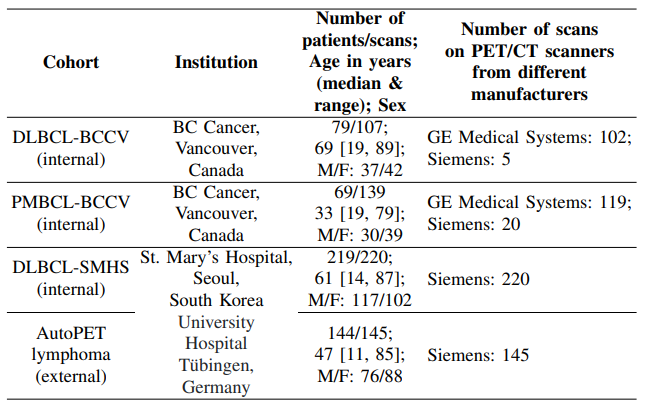
1) Description: In this work, we used a large, diverse and multi-institutional whole-body PET/CT dataset with a total of 611 cases. These scans came from four retrospective cohorts: (i) DLBCL-BCCV: 107 scans from 79 patients with DLBCL from BC Cancer, Vancouver (BCCV), Canada; (ii) PMBCLBCCV: 139 scans from 69 patients with PMBCL from BC Cancer; (iii) DLBCL-SMHS: 220 scans from 219 patients with DLBCL from St. Mary’s Hospital, Seoul (SMHS), South Korea; (iv) AutoPET lymphoma: 145 scans from 144 patients with lymphoma from the University Hospital Tubingen, Germany ¨ [14]. Additional description on the number of scans, patient age and sex, and manufacturers of PET/CT scanner for each cohort is given in Table I. The cohorts (i)-(iii) are collectively referred to as the internal cohort. For cohorts (i) and (ii), ethics approval was granted by the UBC BC Cancer Research Ethics Board (REB) (REB Numbers: H19-01866 and H19-01611 respectively) on 30 Oct 2019 and 1 Aug 2019 respectively. For cohort (iii), the approval was granted by St. Mary’s Hospital, Seoul (REB Number: KC11EISI0293) on 2 May 2011. Due
to the retrospective nature of our data, patient consent was waived for these three cohorts. The cohort (iv) was obtained from the publicly available AutoPET challenge dataset [14] and is referred to as the external cohort.
2) Ground truth annotation: The DLBCL-BCCV, PMBCLBCCV, and DLBCL-SMHS cohorts were separately segmented by three nuclear medicine physicians (referred to as Physician 1, Physician 4, and Physician 5, respectively) from BC Cancer, Vancouver, BC Children’s Hospital, Vancouver, and St. Mary’s Hospital, Seoul, respectively. Additionally, two other nuclear medicine physicians (Physicians 2 and 3) from BC Cancer segmented 9 cases from DLBCL-BCCV cohort which were used for assessing the inter-observer variability (Section IV-D). Physician 4 additionally re-segmented 60 cases from PMBCL-BCCV cohort which were used for assessing the intra-observer variability (Section IV-C). All these expert segmentations were performed using the semi-automatic gradient-based segmentation tool called PETEdge+ from the MIM workstation (MIM software, Ohio, USA).
The AutoPET lymphoma PET/CT data along with their ground truth segmentations were acquired from The Cancer Imaging Archive. These annotations were performed manually by two radiologists from the University Hospital Tubingen, ¨ Germany, and the University Hospital of the LMU, Germany.
B. Networks, tools and code
Four networks were trained in this work, namely, UNet [15], SegResNet [16], DynUNet [17] and SwinUNETR [18]. The former three are 3D CNN-based networks, while SwinUNETR is a transformer-based network. The implementations for these networks were adapted from the MONAI library [19]. The models were trained and validated on Microsoft Azure virtual machine with Ubuntu 16.04, which consisted of 24 CPU cores (448 GiB RAM) and 4 NVIDIA Tesla V100 GPUs (16 GiB RAM each). The code for this work has been open-sourced under the MIT License and can be found in this repository: https://github.com/microsoft/lymphoma-segmentation-dnn.
C. Training methodology
1) Data split: The data from cohorts (i)-(iii) (internal cohort with a total of 466 cases) were randomly split into training (302 scans), validation (76 scans), and internal test (88 scans) sets, while the AutoPET lymphoma cohort (145 scans) was used solely for external testing. The models were first trained on the training set, and the optimal hyperparameters and best models were selected on the validation set. Top models were then tested on the internal and external test sets. Note that the splitting of the internal cohort was performed at the patient level to avoid overfitting the trained model’s parameters to specific patients if their multiple scans happen to be shared between training and validation/test sets.
2) Preprocessing and augmentations: The high-resolution CT images (in the Hounsfield unit (HU)) were down sampled to match the coordinates of their corresponding PET/mask images. The PET intensity values in units of Bq/ml were decay corrected and converted to SUV. During training, we employed a series of non-randomized and randomized transforms to augment the input to the network. The non-randomized transforms included (i) clipping CT intensities in the range of [-154, 325] HU (representing the [3, 97]th quantile of HUs within the lesions across the training and validation sets) followed by min-max normalization, (ii) cropping the region outside the body in PET, CT, and mask images using a 3D bounding box, and (iii) resampling the images to an isotropic voxel spacing of (2.0 mm, 2.0 mm, 2.0 mm) via bilinear interpolation for PET and CT images and nearest-neighbor interpolation for mask images
On the other hand, the randomized transforms were called at the start of every epoch. These included (i) randomly cropping cubic patches of dimensions (N, N, N) from the images, where the cube was centered around a lesion voxel with probability pos/(pos + neg), or around a background voxel with probability neg/(pos + neg), (ii) translations in the range (-10, 10) voxels along all three directions, (iii) axial rotations in the range (−π/15, π/15), and (iv) random scaling by 1.1 in all three directions. We set neg = 1, and pos and N were chosen from the hyperparameter sets {1, 2, 4, 6, 8, 10, 12, 14, 16} and {96, 128, 160, 192, 224, 256} respectively for UNet [20]. After a series of comprehensive ablation experiments, pos = 2 and N = 224 were found to be optimal for UNet. For other networks, pos was set to 2, and the largest N that could be accommodated into GPU memory during training was chosen (since the performance for different values of N were not significantly different from each other, except N = 96 which was significantly worse as compared to other values of N). Hence, SegResNet, DynUNet, and SwinUNETR were trained using N = 192, 160, and 128, respectively. Finally, the augmented PET and CT patches were channel-concatenated to construct the final input to the network.
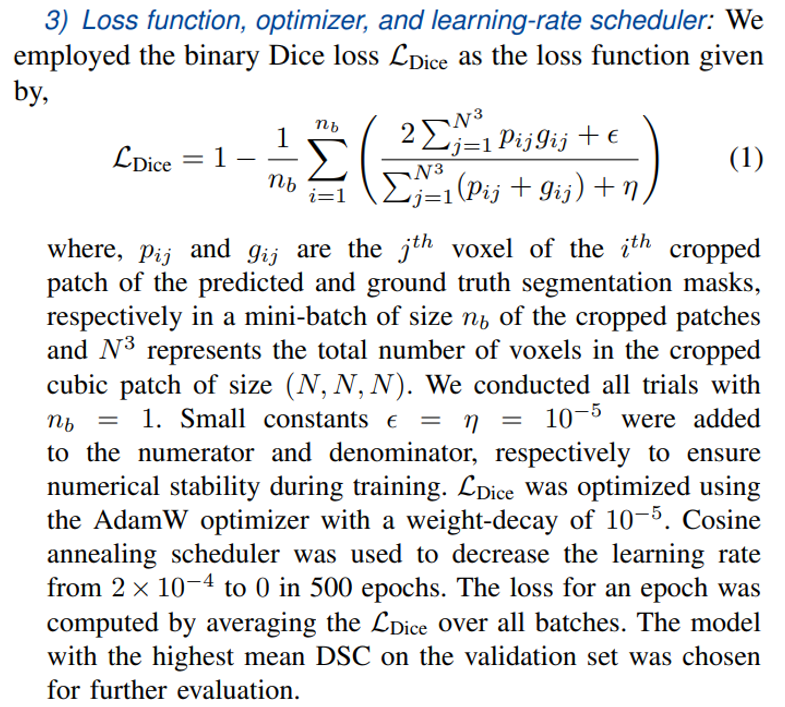

4) Sliding window inference and postprocessing: For the images in the validation/test set, we employed only the nonrandomized transforms. The prediction was made directly on the 2-channel (PET and CT) whole-body images using the sliding-window technique with a cubic window of size (W, W, W), where W was a hyperparameter chosen from the set {96, 128, 160, 192, 224, 256, 288}. The optimal values W were found to 224 for UNet, 192 for SegResNet and DynUnet, and 160 for SwinUNETR. The test set predictions were resampled to the coordinates of the original ground truth masks for computing the evaluation metrics.
D. Evaluation metrics
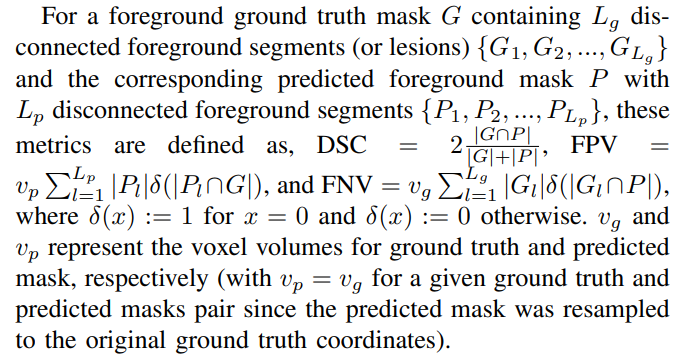
1) Segmentation metrics: To evaluate segmentation performance, we used patient-level foreground DSC, the volumes of false positive connected components that do not overlap with the ground truth foreground (FPV), and the volume of foreground connected components in the ground truth that do not overlap with the predicted segmentation mask (FNV) [14]. We reported the median and interquartile range (IQR) for these metrics on the internal and external test sets. We also report mean DSC with standard deviation on mean. We chose to report the median values since our mean metric values were prone to outliers and our sample median was always higher (lower) for DSC (for FPV and FNV) than the sample mean. An illustration of FPV and FNV is given in Fig. 1 (a).

2) Detection metrics:* Apart from the segmentation metrics discussed above, we also assessed the performance of our models on the test sets via three detection-based metrics for evaluating the detectability of individual lesions within a patient.
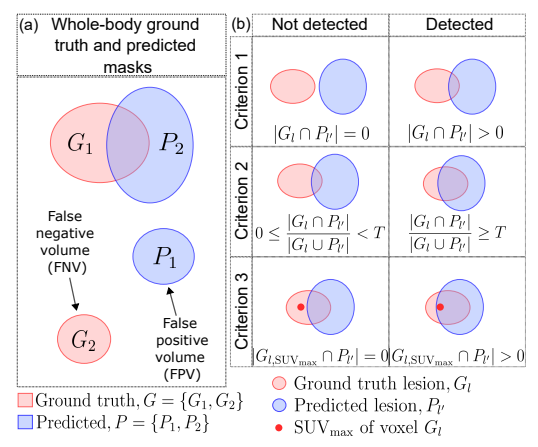
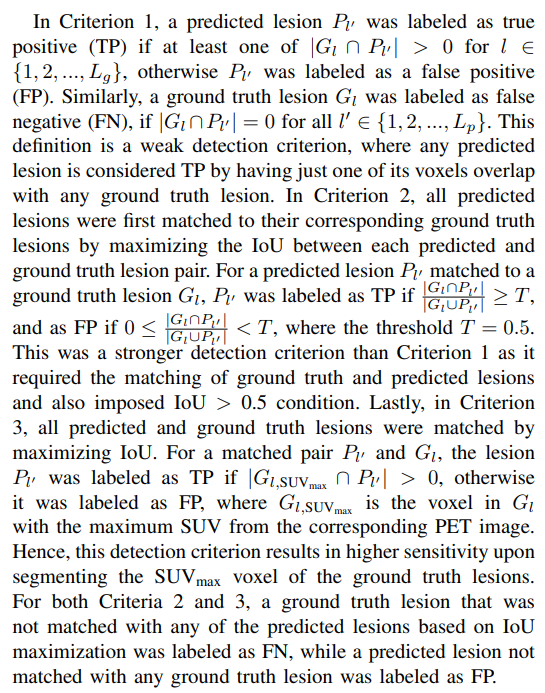


Although the definitions for detection metrics FP and FN might appear similar to the segmentation metrics FPV and FNV, on careful investigation, they are not (Fig. 1 (a) and (b)). FPV and FNV metrics compute the sum of the volumes of all lesions that are predicted in an entirely wrong location (no overlap with ground truth lesions) or lesions that are entirely missed, respectively. Hence, these metrics are defined at the voxel level for each patient. On the other hand, the detection metrics (in Criteria 1, 2, and 3) are defined on a per-lesion basis for each patient.
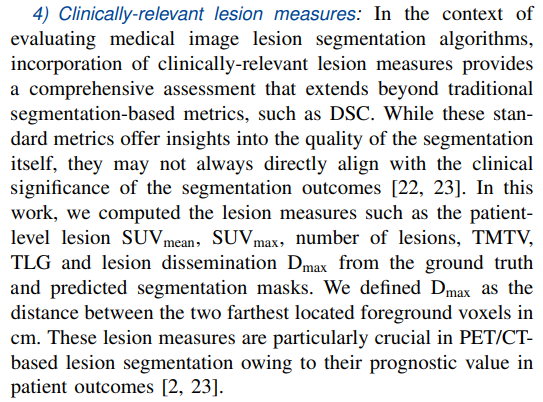


Assessing the reproducibility of these lesion measures enhances the confidence in the segmentation algorithm’s results. Therefore, we conducted paired Student’s t-test analyses to determine the disparity in the means of the distributions between the ground truth and predicted lesion measures (Section IV-A.1). Additionally, similar analyses were carried out to evaluate intra-observer variability, involving two annotations made by the same physician on the same set of cases (Section IV-C).
This paper is